How to use `recipes` package from `tidymodels` for one hot encoding 🛠
Since once of the best way to learn, is to explain, I want to share with you this quick introduction to recipes package, from the tidymodels family.
It can help us to automatize some data preparation tasks.
The overview is:
- How to create a
recipe - How to add a
step - How to do the
prep - Getting the data with
juice! - Apply the prep to new data
- What is the difference between
bakeandjuice? - Dealing with new values in recipes (
step_novel)
Since I'm new to this package, if you have something to add just put in the comments ;)
Introduction
If you are new to R or you do a 1-time analysis, you could not see the main advantage of this, which is -in my opinion- to have most of the data preparation steps in one place. This way is easier to split between dev and prod.
- Dev: The stage in which we create the model
- Prod: The moment in which we run the model with new data
The other big advantage is it follows the tidy philosophy, so many things will be familiar.
How to use recipes for one hot encoding
It is focused on one hot encoding, but many other functions like scaling, applying PCA and others can be performed.
But first, what is one hot encoding?
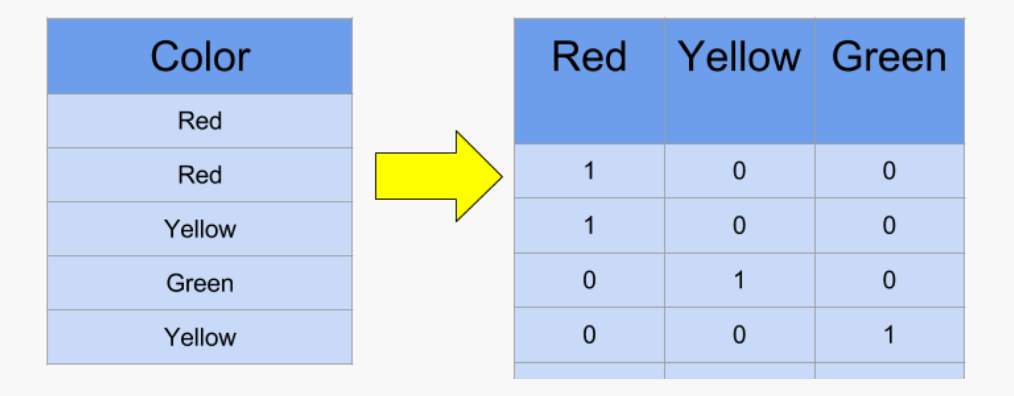
It's a data preparation technique to convert all the categorical variables into numerical, by assigning a value of 1 when the row belongs to the category. If the variable has 100 unique values, the final result will contain 100 columns.
That's why it is a good practice to reduce the cardinality of the variable before continuing Learn more about it in the High Cardinality Variable in Predictive Modeling from the Data Science Live Book 📗.
Let's start the example with recipes!
1st - How to create a recipe
library(recipes)
library(tidyverse)
set.seed(3.1415)
iris_tr=sample_frac(iris, size = 0.7)
rec = recipe( ~ ., data = iris_tr)
rec
## Data Recipe
##
## Inputs:
##
## role #variables
## predictor 5
summary(rec)
## # A tibble: 5 x 4
## variable type role source
## <chr> <chr> <chr> <chr>
## 1 Sepal.Length numeric predictor original
## 2 Sepal.Width numeric predictor original
## 3 Petal.Length numeric predictor original
## 4 Petal.Width numeric predictor original
## 5 Species nominal predictor original
The formula ~ ., specifies that all the variables are predictors (with no outcomes).
Please note now we have two different data types, numeric and nominal (not factor nor character).
2nd - How to add a step
Now we add the step to create the dummy variables, or the one hot encoding, which can be seen as the same.
When we do the one hot encoding (one_hot = T), all the levels will be present in the final result. Conversely, when we create the dummy variables, we could have all of the variables, or one less (to avoid the multi-correlation issue).
rec_2 = rec %>% step_dummy(Species, one_hot = T)
rec_2
## Data Recipe
##
## Inputs:
##
## role #variables
## predictor 5
##
## Operations:
##
## Dummy variables from Species
Now we see the dummy step.
3rd - How to do the prep
prep is like putting all the ingredients together, but we didn't cook yet!
It generates the metadata to do the data preparation.
As we can see here:
# Aplico la receta, que tiene 1 step, a los datos
d_prep=rec_2 %>% prep(training = iris_tr, retain = T)
d_prep
## Data Recipe
##
## Inputs:
##
## role #variables
## predictor 5
##
## Training data contained 105 data points and no missing data.
##
## Operations:
##
## Dummy variables from Species [trained]
Note we are in the "training" or dev stage. That's why we see the parameter training.
We will see retain = T in the next step.
Checking:
summary(d_prep)
## # A tibble: 7 x 4
## variable type role source
## <chr> <chr> <chr> <chr>
## 1 Sepal.Length numeric predictor original
## 2 Sepal.Width numeric predictor original
## 3 Petal.Length numeric predictor original
## 4 Petal.Width numeric predictor original
## 5 Species_setosa numeric predictor derived
## 6 Species_versicolor numeric predictor derived
## 7 Species_virginica numeric predictor derived
Whoila! 🎉 We have the 3-new derived columns (one hot), and it removed the original Species.
4th - Getting the data with juice!
Using juice function:
d2=juice(d_prep)
head(d2)
## # A tibble: 6 x 7
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species_setosa
## <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 5 3 1.6 0.2 1
## 2 6.9 3.2 5.7 2.3 0
## 3 6.3 3.3 4.7 1.6 0
## 4 5.3 3.7 1.5 0.2 1
## 5 6.3 2.3 4.4 1.3 0
## 6 6.7 3 5.2 2.3 0
## # … with 2 more variables: Species_versicolor <dbl>,
## # Species_virginica <dbl>
juice worked because we retained the training data in the 3rd step (retain = T). Otherwise it would have returned:
⚠️ Error: Use retain = TRUE in prep to be able to extract the training set
5th - Apply the prep to new data
Now imagine we have new data as follows:
iris_new=sample_n(iris, size = 5) # taking 5 random rows
d_baked=bake(d_prep, new_data = iris_new)
d_baked
## # A tibble: 5 x 7
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species_setosa
## <dbl> <dbl> <dbl> <dbl> <dbl>
## 1 6.4 3.2 4.5 1.5 0
## 2 4.6 3.4 1.4 0.3 1
## 3 5.2 2.7 3.9 1.4 0
## 4 4.8 3.4 1.6 0.2 1
## 5 4.8 3 1.4 0.3 1
## # … with 2 more variables: Species_versicolor <dbl>,
## # Species_virginica <dbl>
It worked!
bake receives the prep object (d_prep) and it applies to the new_data (iris_new)
What is the difference between bake and juice?
From this perspective given the training data, following data frames are the same:
d_tr_1=bake(d_prep, new_data = iris_tr)
d_tr_2=d2=juice(d_prep) # with retain=T
identical(d_tr_1, d_tr_2)
## [1] TRUE
Dealing with new values in recipes
Simulate a new value:
new_row=iris[1,] %>% mutate(Species=as.character(Species))
new_row[1, "Species"]="i will break your code"
new_row
## Sepal.Length Sepal.Width Petal.Length Petal.Width Species
## 1 5.1 3.5 1.4 0.2 i will break your code
We use bake to convert the new data set:
d2_b=bake(d_prep, new_data = new_row)
## Warning: There are new levels in a factor: i will break your code
The solution! Use step_novel
(Thanks to Max Kuhn)
When we do the prep, we have to add step_novel. So any new value will be assigned to the _new category.
We will start right from the beginning:
rec_2_bis = recipe( ~ ., data = iris_tr) %>%
step_novel(Species) %>%
step_dummy(Species, one_hot = T)
prep_bis = prep(rec_2_bis, training = iris_tr)
Get to final data, and check it:
processed = bake(prep_bis, iris_tr)
funModeling::df_status(processed)
## variable q_zeros p_zeros q_na p_na q_inf p_inf type unique
## 1 Sepal.Length 0 0.00 0 0 0 0 numeric 32
## 2 Sepal.Width 0 0.00 0 0 0 0 numeric 23
## 3 Petal.Length 0 0.00 0 0 0 0 numeric 42
## 4 Petal.Width 0 0.00 0 0 0 0 numeric 20
## 5 Species_setosa 68 64.76 0 0 0 0 numeric 2
## 6 Species_versicolor 69 65.71 0 0 0 0 numeric 2
## 7 Species_virginica 73 69.52 0 0 0 0 numeric 2
## 8 Species_new 105 100.00 0 0 0 0 numeric 1
Please note that Species_new has been automatically created (with zeros).
👉 This ensures it runs well once in production.
Now let's see what happen when we have the new value:
new_row_2=bake(prep_bis, new_data = new_row)
new_row_2 %>% select(Species_new)
## # A tibble: 1 x 1
## Species_new
## <dbl>
## 1 1
It works!
Conclusions 💡
The recipes package seems to be a good way to standardize certain data preparation tasks.
Probably one of the strongest points in R, alongside the dplyr package.
📌 Take care of the data pipeline, it is what interviewers will ask you for.
I tried to cover with simple and reproducible examples, many of the situations that happen when we work with productive environments, in the Data Science Live Book 📗 (open-source).
Have fun 🚀
📬 You can found me at: Linkedin & Twitter.
References:
- Basic recipes example
- Modeling with parsnip and tidymodels by Benjamin Sorensen.
- Creating and Preprocessing a Design Matrix with Recipes (video)